User Survey:
In order to improve our services, please provide your feedback in our LipidCreator Survey.
Citing LipidCreator
Please reference LipidCreator by citing the following publication:
Peng, B., Kopczynski, D., Pratt, B.S. et al. LipidCreator workbench to probe the lipidomic landscape. Nat Commun 11, 2057 (2020). https://doi.org/10.1038/s41467-020-15960-z
LipidCreator Workshop Material Skyline Tool Store Source Code
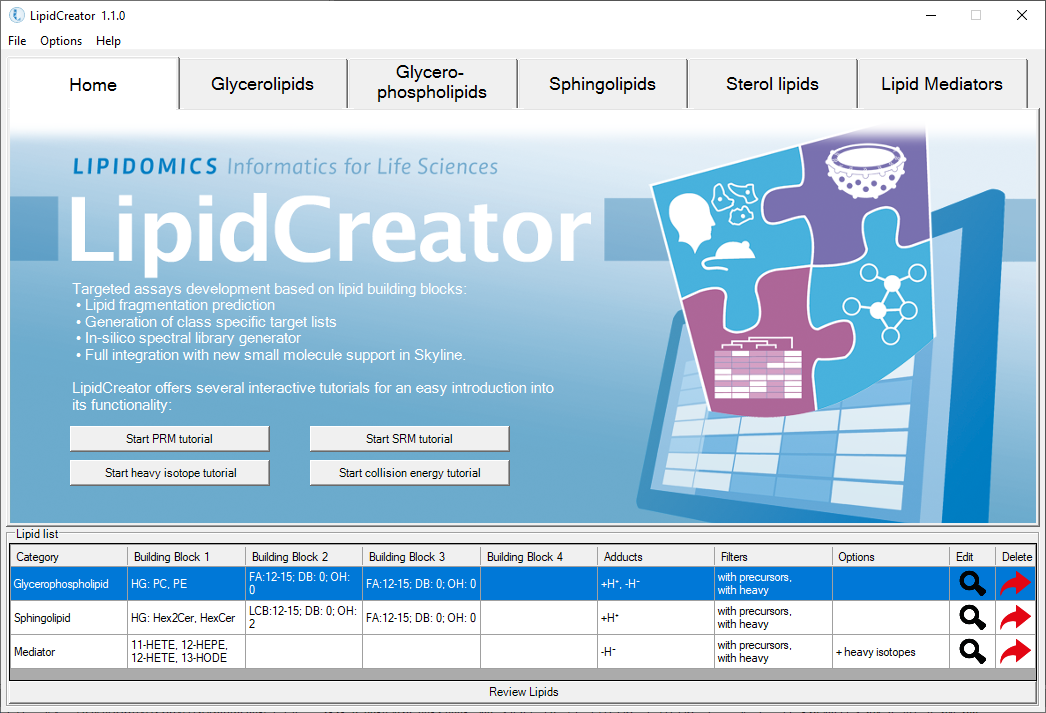
LipidCreator is a lipid building block-based workbench and knowledge base for the semi-automatic generation of targeted lipidomics mass spectrometry assays and in silico spectral libraries.
- Lipid Categories: The following lipid categories are currently supported: sphingolipids (SL), glycerolipids (GL), glycerophospholipids (PL), cholesterol and its derivates (Ch, ChE), as well as fatty acyl mediators (LM).
- SRM / PRM assay generation: LipidCreator can calculate masses and transitions for both unlabeled and labeled lipid species and their derived fragment ions, covering over 60 lipid classes and a lipid array of 1012 lipid molecules.
- Lipid Nomenclature and Translation: LipidCreator uses the consensus nomenclature recommended by the lipidomics standards initiative and provides a lipid nomenclature translator that supports LIPID MAPS names.
- Visual Inspection: LipidCreator further contains a visual inspection level for lipid and fragment structures.
- In Silico Spectral Library & Collision Energy Optimization: Besides the computation of fragment ions, another crucial feature to generate an in silico spectral library is the ability to determine the relative intensities of fragment ions at different defined collision energies (CE). We therefore trained non-linear regression models on empirical data from the measurements of lipid mediator standards on Thermo QExactive HF and Agilent 6545 Q-TOF instruments.
- Skyline Plugin: LipidCreator can work standalone but it is also fully integrable into Skyline and its small molecule support, allowing vendor-independent assay development and optimization, data visualization, peak integration and identification, as well as quality control of MS and MS/MS lipidomics data.
LipidCreator can be installed directly from the Skyline Tool Store. - GUI or CLI: LipidCreator can be used with a user-friendly graphical user interface (GUI) or via the command line interface (CLI).
- Performance: Using the batch-processing mode, LipidCreator computes precursor-product ion pairs at a rate of 60,000 pairs per second on a standard notebook (i5-4310M CPU @ 2.70GHz with 8GB main memory).
- Workflow Integration: It can be integrated into KNIME and Galaxy workflows as a native node via its command line interface on Linux and Windows.
In combination with Skyline, LipidCreator provides an integrated and user-friendly targeted workflow development environment for comprehensive lipidomics analyses.
Please see the other articles below for more information on how to install and use LipidCreator.
LipidCreator Release History
Contact and Help Desk
For any further information, please contact us.
Documentation
LipidCreator comes with four interactive tutorials and a manual that is accessible once you start the application. Please consult the release notes below for installation instructions and additional training / workshop material.
